Sayan Goswami, assistant professor of computer science at St. Mary’s College of Maryland, recently co-authored a pioneering study in the journal Bioinformatics titled "Rawsamble: Overlapping Raw Nanopore Signals using a Hash-based Seeding Mechanism." The research introduces a novel method for analyzing genetic data that drastically reduces the time and computing power required for genome assembly.
In modern genomics, Nanopore sequencing works by passing DNA strands through a microscopic pore and measuring the resulting electrical current. Usually, these "raw" signals must undergo a computationally intensive translation process called "basecalling"- converting electrical squiggles into the familiar DNA bases (A, C, G, and T) before scientists can begin piecing the genome together.
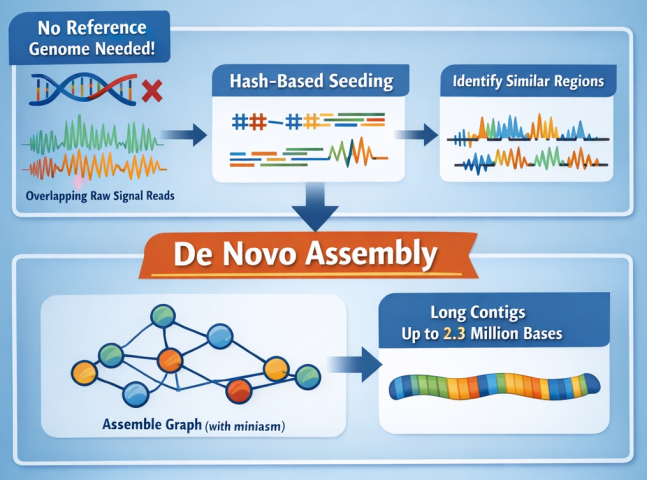
Goswami and an international team of collaborators developed Rawsamble, a mechanism that allows researchers to skip the basecalling step entirely. By using a "hash-based" search to identify similarities between raw electrical signals directly, the team proved that genomes could be assembled with significantly greater speed and a much smaller memory footprint than traditional methods.
"Our goal was to enable the direct analysis of raw signals without needing a reference genome," the researchers noted. The study’s evaluations showed that Rawsamble can be up to ten times faster than current state-of-the-art tools while maintaining high accuracy. This breakthrough suggests a future where complex genetic analysis can be performed on more accessible, less expensive hardware, democratizing high-level research.